Service Charges and Ordering
Service Charges
Effective 1-Sep-2023
| Assay Type | Maximum Number of Samples per Chip | HKU Academic Research | Other Academic Research | ||
| per Sample | per Chip | per Sample | per Chip | ||
| RNA 6000 Nano | 12 | HK$ 67 | HK$ 800 | HK$ 84 | HK$ 1,000 |
| DNA High Sensitivity | 11 | HK$ 92 | HK$ 1,010 | – | HK$ 1,265 |
For commercial clients, please enquire.
*IMPORTANT*: To qualify for HKU pricing, the investigator must be a regular employee of HKU AND payment must be made from an internal HKU financial account that qualifies for internal transfer. Overhead charge is required for all incoming external funding.
Service Ordering
Please submit service request through iLab.
Workflow
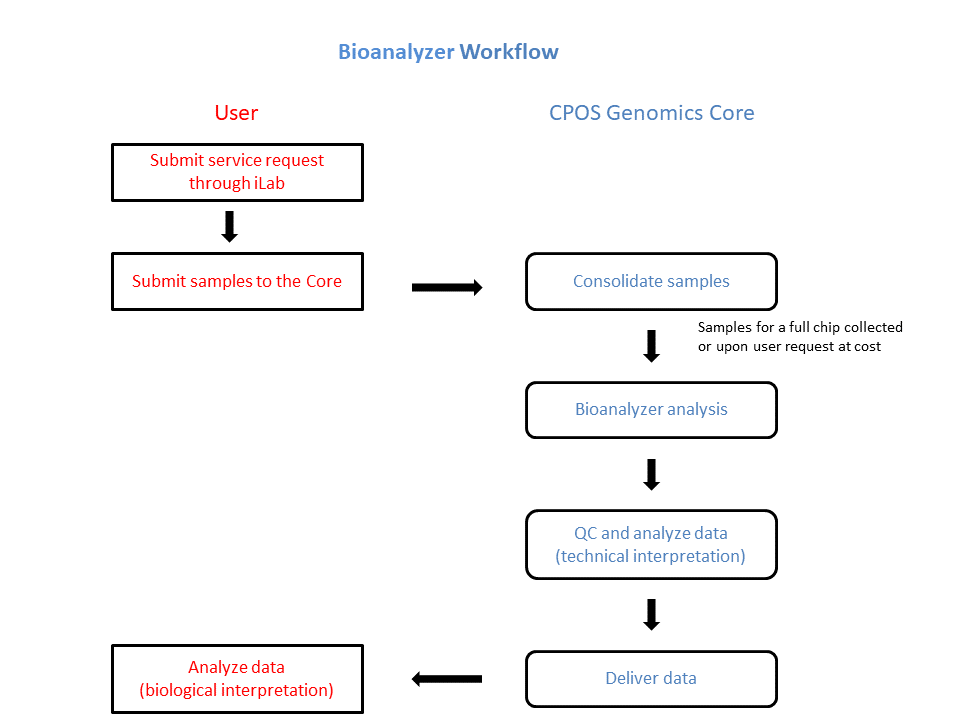
Sample Preparation and Submission
The Core only keeps a small amount of stock of the RNA 6000 Nano chip and High Sensitivity DNA chip. For requests for more than 5 chips and/or for assays not listed below, please contact platform specialists for further information and arrangement.
Sample Preparation and Submission Guidelines
(1) Prepare samples in 0.5 mL tubes according to the sensitivity range and buffer concentration provided in below table. Based on our in-house experience, it is advisable to aim for a concentration around the midpoint of the sensitivity range, as this tends to yield the best results. Concentrations towards the lower end of the range may not provide reliable data.
| Bioanalyzer Assay | Sizing Range | Recommended Concentration Range | Maximum Buffer Concentration | Minimum Sample Volume | Number of Samples per Chip |
| RNA 6000 Nano total RNA# | 25 – 6000 nt | 30 – 500 ng/μL | 100 mM Tris 0.1 mM EDTA or 125 mM NaCl 15 mM MgCl2 | 3 μL | 12 |
RNA 6000 Nano mRNA#| 25 – 6000 nt | 30 – 250 ng/μL | 100 mM Tris | 0.1 mM EDTA or 125 mM NaCl 15 mM MgCl2 3 μL | 12 | |
| High Sensitivity DNA | 50 – 7000 bp | Fragment: 0.1 – 1 ng/μL * Smear: 2 – 10 ng/μL ** | 10 mM Tris and 1 mM EDTA | 3 μL | 11 |
#Users are required to commit service for a whole chip with any customized run setup such as RNA 6000 Nano Assays for Prokaryotic RNA run, RNA 6000 Nano mRNA Assay, and others.
* Samples with a single peak, e.g. amplicon
** Samples with a broad peak, e.g. NGS library
(2) Clearly label each tube with sample ID.
(3) Submit service request through iLab. Smear analysis from 200 bp to 1,000 bp would be provided for High Sensitivity DNA assay. Please specify if other ranges are needed.
(4) Submit completed Bioanalyzer sample submission form as attachment in iLab.
(5) Contact platform specialist for sample submission appointment.
(6) Bring the samples to the Bioanalyzer unit at the Core.
Special notes for Illumina Sequencing (Next Generation Sequencing) users
Please contact our Illumina platform specialists (nextgenseq.cpos@hku.hk) prior to submitting your samples to the Core.
*IMPORTANT*: It is not recommended to rely on the concentration indicated on the Bioanalyzer report for NGS library quantity check and NGS sequencing preparation. We advise using qPCR for accurate measurement of NGS library concentration.
For more details about submission requirements for Illumina Sequencing, kindly visit the Illumina Sequencing Sample Requirements and Submission webpage.
If your samples do not meet above requirements, please feel free to reach out to us (bioanalyzer.cpos@hku.hk / 2831-5477) for assistance.
Data Collection
Processing Time
The expected turnaround time for a full Bioanalyzer chip analysis is usually 1-2 working days. Requests with sample number less than a full chip will be put into a job queue until enough samples are consolidated for a full chip analysis. As such, the turnaround time will vary depending on the number of samples collected.
Data Delivery
Results will be delivered in PDF format via our Online Data Delivery System (ODDS).
Unless otherwise specified, a smear analysis between 200bp to 1,000bp would be included in the report for all DNA assays.
Terms of Service
Leftover Samples
Leftover samples will be kept for one week after the delivery of results.
Users are welcome to collect any leftover samples during this period, after which the samples will be discarded without further notice.
Please contact our platform specialist for sample pick-up arrangement.
Contact
Mr LAM, Jason




